Processing pipeline details¶
fmriprep adapts its pipeline depending on what data and metadata is
available is used as the input. For example, slice timing correction will be
performed only if the SliceTiming metadata field is found for the input
dataset.
High-level view of the basic pipeline (for single-band datasets, without slice-timing information and no fieldmap acquisitions):
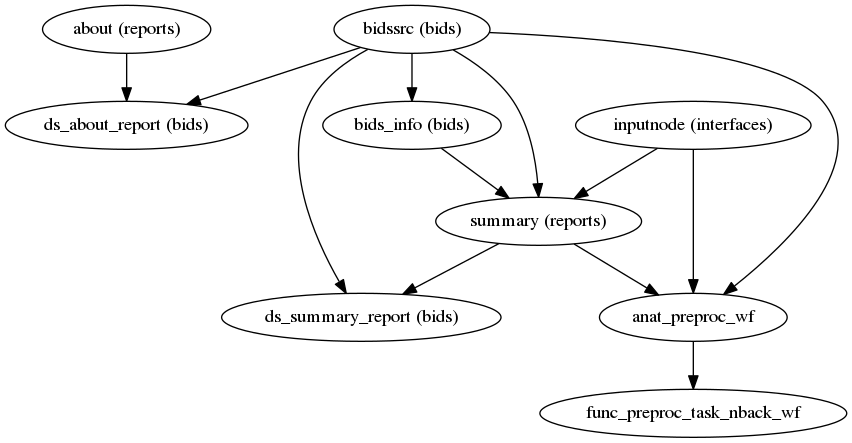
(Source code, png, svg, pdf)
T1w/T2w preprocessing¶
fmriprep.workflows.anatomical.init_anat_preproc_wf
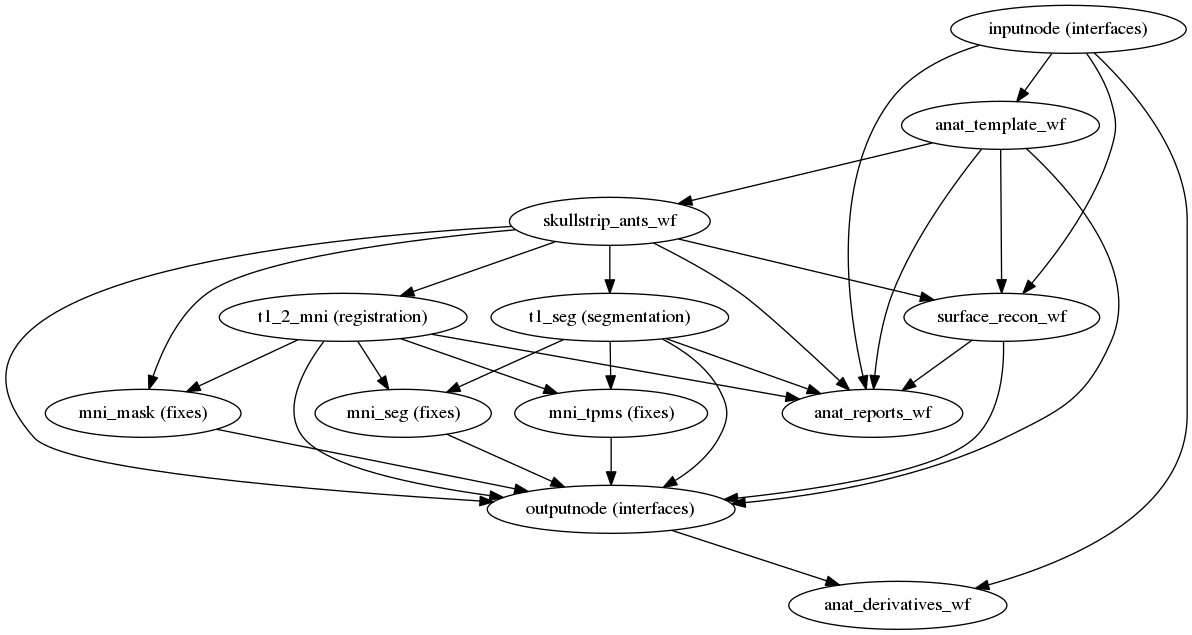
(Source code, png, svg, pdf)
The anatomical sub-workflow begins by constructing a template image by conforming any T1-weighted images to RAS orientation and a common voxel size, and, in the case of multiple images, merges them into a single template (see Longitudinal processing). This template is then skull-stripped, and the white matter/gray matter/cerebrospinal fluid segments are found. Finally, a non-linear registration to the MNI template space is estimated.
Longitudinal processing¶
In the case of multiple sessions, T1w images are merged into a single template
image using FreeSurfer’s mri_robust_template.
This template may be unbiased, or equidistant from all source images, or
aligned to the first image (determined lexicographically by session label).
For two images, the additional cost of estimating an unbiased template is
trivial and is the default behavior, but, for greater than two images, the cost
can be a slowdown of an order of magnitude.
Therefore, in the case of three or more images, fmriprep constructs
templates aligned to the first image, unless passed the --longitudinal
flag, which forces the estimation of an unbiased template.
Note
The preprocessed T1w image defines the T1w space.
In the case of multiple T1w images, this space may not be precisely aligned
with any of the original images.
Reconstructed surfaces and functional datasets will be registered to the
T1w space, and not to the input images.
Surface preprocessing¶
fmriprep.workflows.anatomical.init_surface_recon_wf
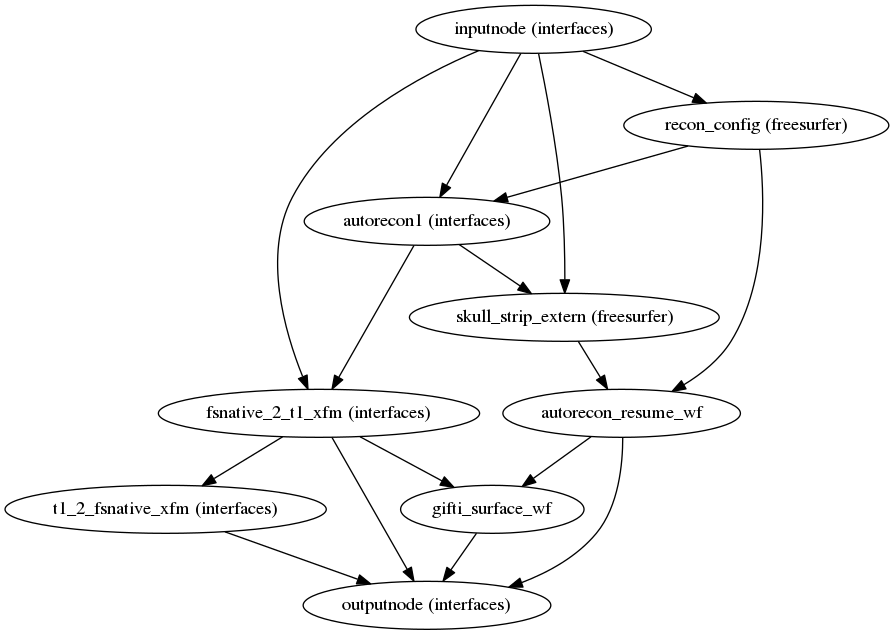
(Source code, png, svg, pdf)
fmriprep uses FreeSurfer to reconstruct surfaces from T1w/T2w
structural images.
If enabled, several steps in the fmriprep pipeline are added or replaced.
All surface preprocessing may be disabled with the --no-freesurfer flag.
If FreeSurfer reconstruction is performed, the reconstructed subject is placed in
<output dir>/freesurfer/sub-<subject_label>/ (see FreeSurfer Derivatives).
Surface reconstruction is performed in three phases.
The first phase initializes the subject with T1w and T2w (if available)
structural images and performs basic reconstruction (autorecon1) with the
exception of skull-stripping.
For example, a subject with only one session with T1w and T2w images
would be processed by the following command:
$ recon-all -sd <output dir>/freesurfer -subjid sub-<subject_label> \
-i <bids-root>/sub-<subject_label>/anat/sub-<subject_label>_T1w.nii.gz \
-T2 <bids-root>/sub-<subject_label>/anat/sub-<subject_label>_T2w.nii.gz \
-autorecon1 \
-noskullstrip
The second phase imports the brainmask calculated in the T1w/T2w preprocessing
sub-workflow.
The final phase resumes reconstruction, using the T2w image to assist
in finding the pial surface, if available.
See init_autorecon_resume_wf() for
details.
Reconstructed white and pial surfaces are included in the report.
If T1w voxel sizes are less than 1mm in all dimensions (rounding to nearest
.1mm), submillimeter reconstruction is used, unless disabled with
--no-submm-recon.
In order to bypass reconstruction in fmriprep, place existing reconstructed
subjects in <output dir>/freesurfer prior to the run.
fmriprep will perform any missing recon-all steps, but will not perform
any steps whose outputs already exist.
lh.midthickness and rh.midthickness surfaces are created in the subject
surf/ directory, corresponding to the surface half-way between the gray/white
boundary and the pial surface.
The smoothwm, midthickness, pial and inflated surfaces are also
converted to GIFTI format and adjusted to be compatible with multiple software
packages, including FreeSurfer and the Connectome Workbench.
BOLD preprocessing¶
fmriprep.workflows.bold.base.init_func_preproc_wf
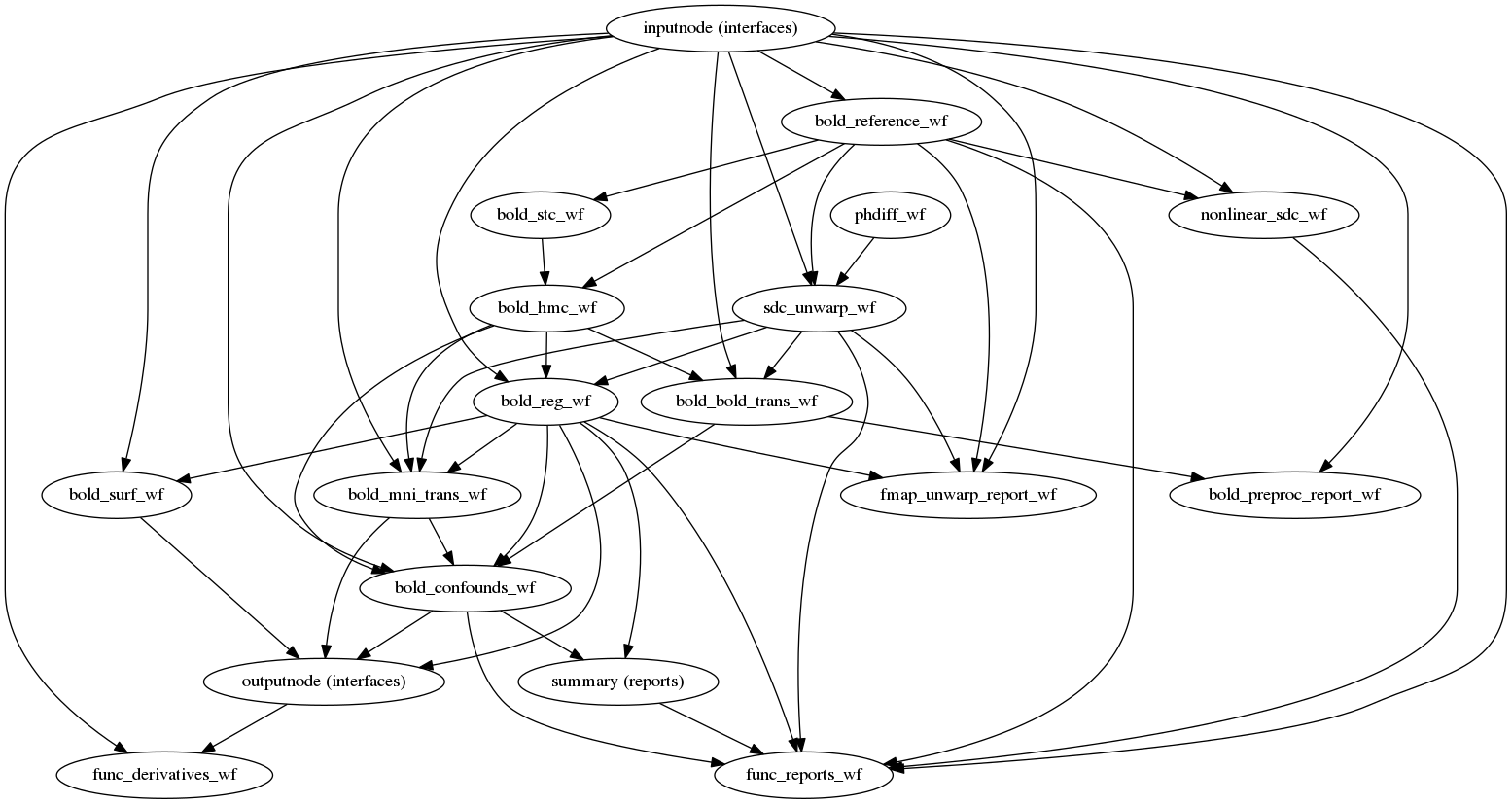
(Source code, png, svg, pdf)
Preprocessing of BOLD files is split into multiple sub-workflows decribed below.
BOLD reference image estimation¶
fmriprep.workflows.bold.util.init_bold_reference_wf
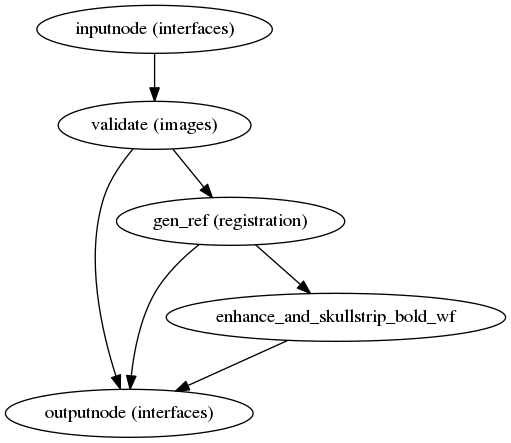
(Source code, png, svg, pdf)
This workflow estimates a reference image for a BOLD series, which is used to estimate head motion and register BOLD series to T1, and performs skull-stripping and contrast enhancement. If T1-saturation effects (“dummy scans” or non-steady state volumes) are detected they are used as reference due to their superior tissue contrast. Otherwise a median of motion corrected subset of volumes is used.
Skullstripping of the reference image is performed using Nilearn.
Slice time correction¶
fmriprep.workflows.bold.stc.init_bold_stc_wf
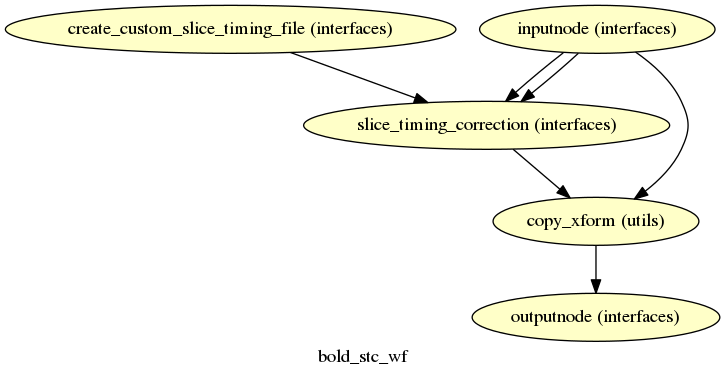
(Source code, png, svg, pdf)
If the SliceTiming field is present in the input dataset metadata, this
workflow will be included to perform slice time correction prior to head motion
estimation.
Slice time correction is performed using AFNI 3dTShift.
All slices are realigned in time to the middle of each TR.
Slice time correction can be disabled with --ignore slicetiming command
line argument.
If a BOLD series has fewer than 5 usable (steady-state) volumes, slice time
correction will be disabled.
Head-motion estimation¶
fmriprep.workflows.bold.hmc.init_bold_hmc_wf
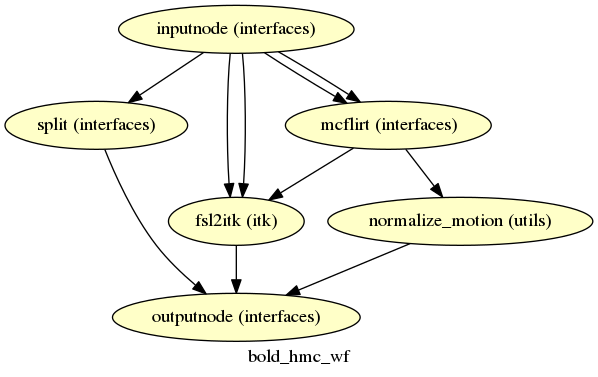
(Source code, png, svg, pdf)
FSL MCFLIRT is used to estimate motion transformations using an automatically estimated reference scan (see BOLD reference image estimation).
Susceptibility Distortion Correction (SDC)¶
One of the major problems that affects EPI data is the spatial distortion caused by the inhomogeneity of the field inside the scanner. The are four broad families of methodologies for mapping the field:
- Phase-difference techniques: to estimate the fieldmap, these methods measure the phase evolution in time between two close GRE acquisitions. Corresponds to the sections 8.9.1 and 8.9.2 of the BIDS specification.
- Direct field-mapping: some sequences (such as SE) are able to measure the fieldmap directly. Corresponds to section 8.9.3 of BIDS.
- Blip-up/blip-down or phase encoding polarity: pepolar: techniques acquire at least two images distorted due to the inhomogeneity of the field, but varying in PE direction. Corresponds to 8.9.4 of BIDS.
- Point-spread function acquisition: Not supported by FMRIPREP.
Once the field-map is estimated, the distortion can be accounted for. Fieldmap processing in FMRIPREP is structured as follows:
- Identifying field-mapping resources: the input BIDS dataset is queried to find the available field-mapping techniques and the appropriate processing workflows are set-up (applies to phase-difference and direct field-mapping techniques).
- Fieldmap estimation: all the estimation workflows produce a displacement field ready to be used in the correction step (applies to phase-difference and direct field-mapping techniques).
- Unwarping EPIs: the correction step is applied (for phase encoding polarity this step also involves distortion correction displacement field estimation).
If the dataset metadata indicate tha more than one field map acquisition is
IntendedFor (see BIDS Specification section 8.9) the following priority will
be used:
- Blip-up/blip-down
- Direct field-mapping
- Phase-difference techniques
Additionally, FMRIPREP now experimentally supports displacement field estimation in the absence of fieldmaps. See Fieldmap-less estimation (experimental) for further details.
Pre-processed BOLD in native space¶
fmriprep.workflows.bold.resampling.init_bold_preproc_trans_wf
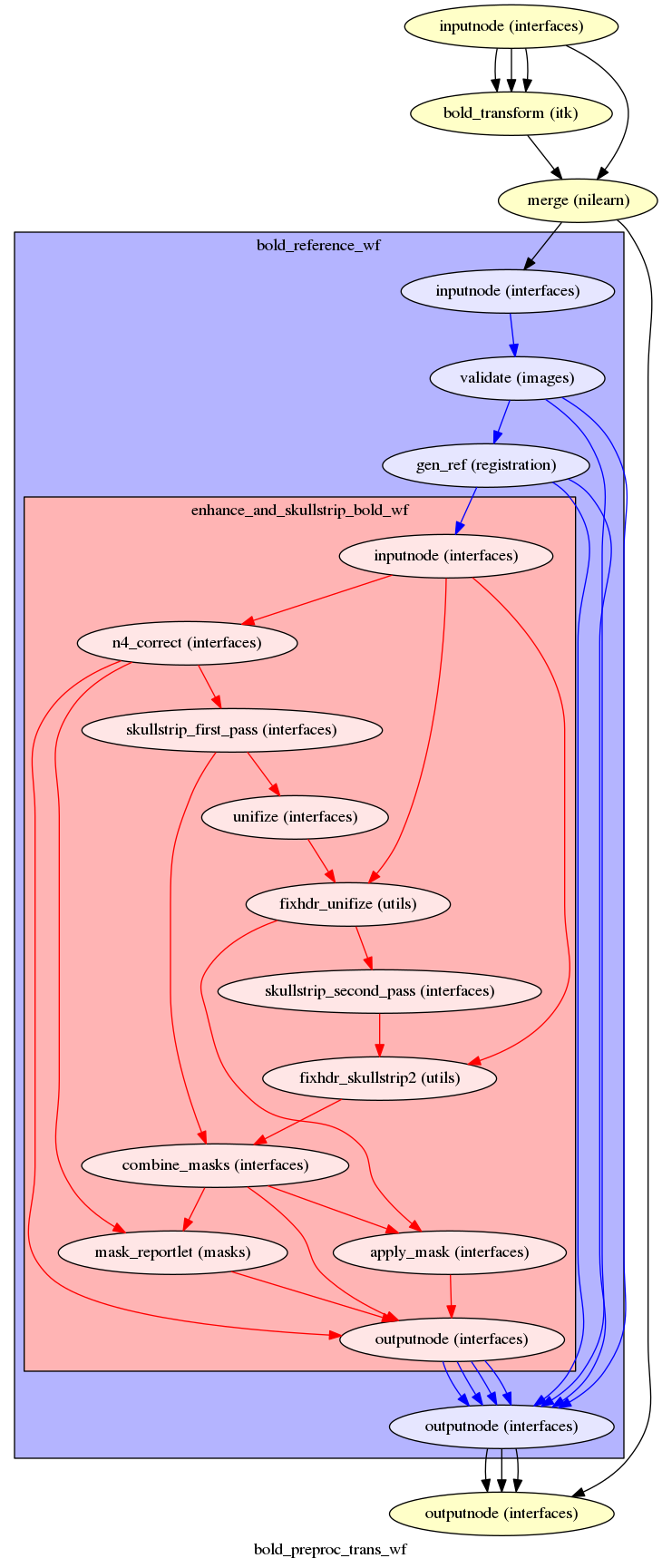
(Source code, png, svg, pdf)
EPI to T1w registration¶
fmriprep.workflows.bold.registration.init_bold_reg_wf
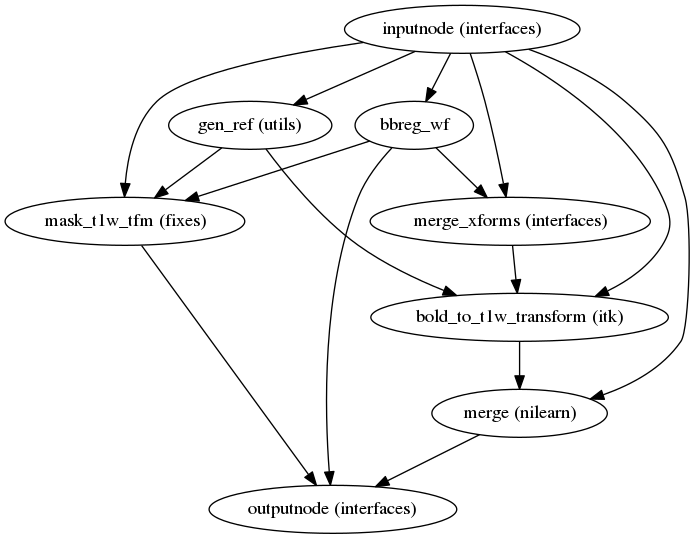
(Source code, png, svg, pdf)
The reference EPI image of each run is aligned by the bbregister routine to the
reconstructed subject using the gray/white matter boundary (FreeSurfer’s
?h.white surfaces).
If FreeSurfer processing is disabled, FLIRT is performed with the BBR cost function, using the FAST segmentation to establish the gray/white matter boundary.
EPI to MNI transformation¶
fmriprep.workflows.bold.resampling.init_bold_mni_trans_wf
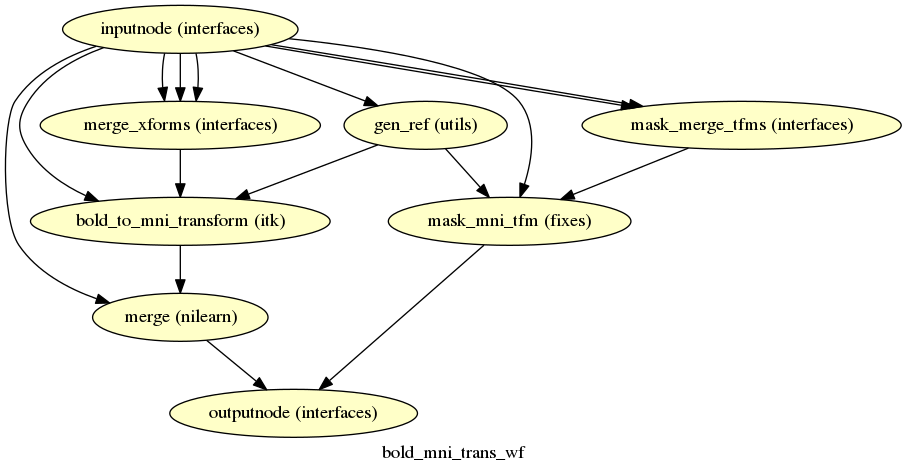
(Source code, png, svg, pdf)
This sub-workflow concatenates the transforms calculated upstream (see Head-motion estimation, Susceptibility Distortion Correction (SDC) (if fieldmaps are available), EPI to T1w registration, and a T1w-to-MNI transform from T1w/T2w preprocessing) to map the EPI image to standardized MNI space. It also maps the T1w-based mask to MNI space.
Transforms are concatenated and applied all at once, with one interpolation (Lanczos) step, so as little information is lost as possible.
EPI sampled to FreeSurfer surfaces¶
fmriprep.workflows.bold.resampling.init_bold_surf_wf
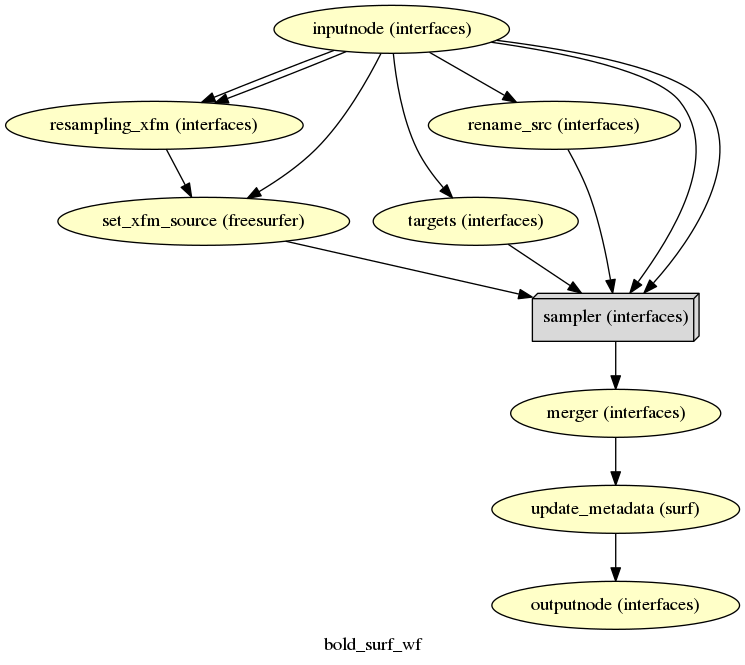
(Source code, png, svg, pdf)
If FreeSurfer processing is enabled, the motion-corrected functional series (after single shot resampling to T1w space) is sampled to the surface by averaging across the cortical ribbon. Specifically, at each vertex, the segment normal to the white-matter surface, extending to the pial surface, is sampled at 6 intervals and averaged.
Surfaces are generated for the “subject native” surface, as well as transformed to the
fsaverage template space.
All surface outputs are in GIFTI format.
Confounds estimation¶
fmriprep.workflows.bold.confounds.init_bold_confs_wf
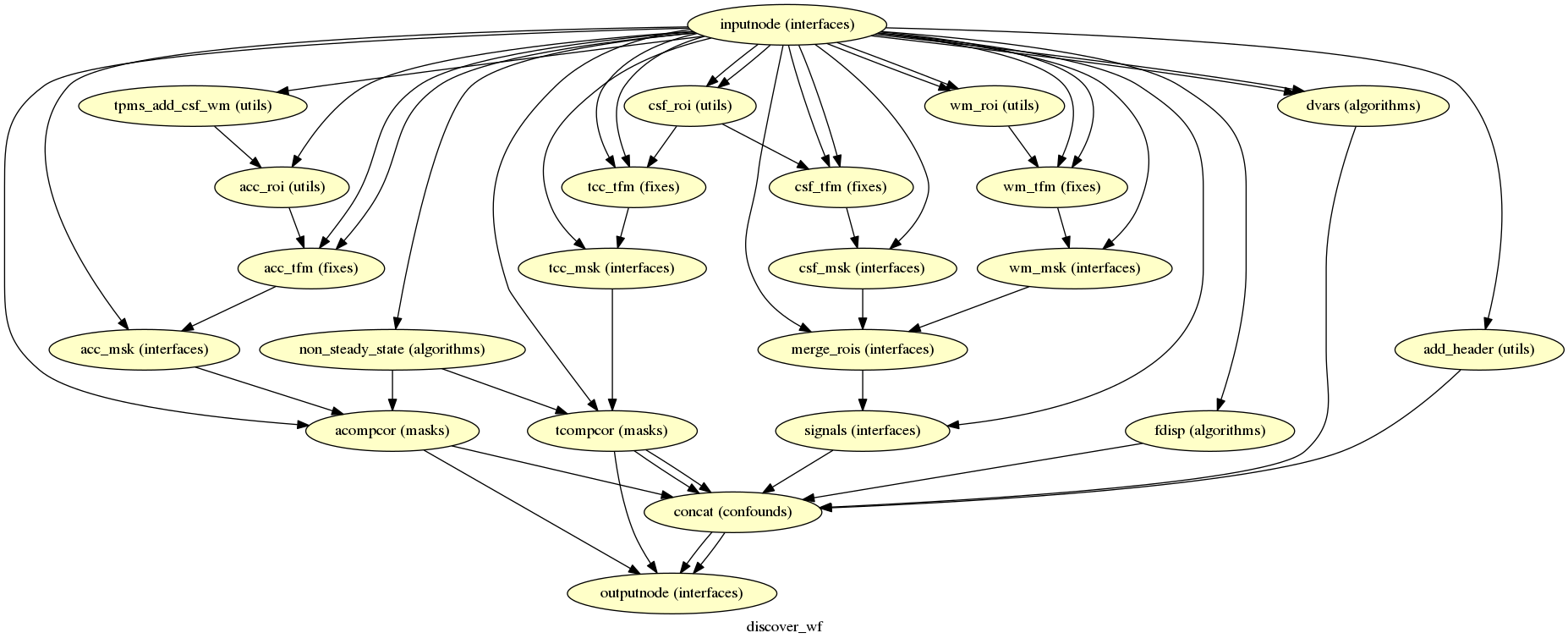
(Source code, png, svg, pdf)
Given a motion-corrected fMRI, a brain mask, MCFLIRT movement parameters and a segmentation, the discover_wf sub-workflow calculates potential confounds per volume.
Calculated confounds include the mean global signal, mean tissue class signal,
tCompCor, aCompCor, Framewise Displacement, 6 motion parameters, DVARS, and, if
the --use-aroma flag is enabled, the noise components identified by ICA-AROMA
(those to be removed by the “aggressive” denoising strategy).
Note: non-aggressive AROMA denoising is a fundamentally different procedure from its “aggressive” counterpart and cannot be performed only by using a set of noise regressors (a separate GLM with both noise and signal regressors needs to be used). Therefore instead of regressors FMRIPREP produces non-aggressive denoised 4D NIFTI files in the MNI space:
*bold_space-MNI152NLin2009cAsym_variant-smoothAROMAnonaggr_brainmask.nii.gz
Additionally, the MELODIC mix and noise component indices will
be generated, so non-aggressive denoising can be manually performed in the T1w space with fsl_regfilt, e.g.:
fsl_regfilt -i sub-<subject_label>_task-<task_id>_bold_space-T1w_preproc.nii.gz \
-f $(cat sub-<subject_label>_task-<task_id>_bold_AROMAnoiseICs.csv) \
-d sub-<subject_label>_task-<task_id>_bold_MELODICmix.tsv \
-o sub-<subject_label>_task-<task_id>_bold_space-<space>_AromaNonAggressiveDenoised.nii.gz
A visualisation of the AROMA component classification is also included in the HTML reports.