Unwarping EPIs¶
Apply susceptibility distortion correction (SDC)¶
Abbreviations
- fmap
- fieldmap
- VSM
- voxel-shift map – a 3D nifti where displacements are in pixels (not mm)
- DFM
- displacements field map – a nifti warp file compatible with ANTs (mm)
-
fmriprep.workflows.fieldmap.unwarp.init_pepolar_unwarp_wf(fmaps, bold_file, omp_nthreads, layout=None, fmaps_pes=None, bold_file_pe=None, name='pepolar_unwarp_wf')[source]¶ This workflow takes in a set of EPI files with opposite phase encoding direction than the target file and calculates a displacements field (in other words, an ANTs-compatible warp file).
This procedure works if there is only one ‘_epi’ file is present (as long as it has the opposite phase encoding direction to the target file). The target file will be used to estimate the field distortion. However, if there is another ‘_epi’ file present with a matching phase encoding direction to the target it will be used instead.
Currently, different phase encoding dimension in the target file and the ‘_epi’ file(s) (for example ‘i’ and ‘j’) is not supported.
The warp field correcting for the distortions is estimated using AFNI’s 3dQwarp, with displacement estimation limited to the target file phase encoding direction.
It also calculates a new mask for the input dataset that takes into account the distortions.
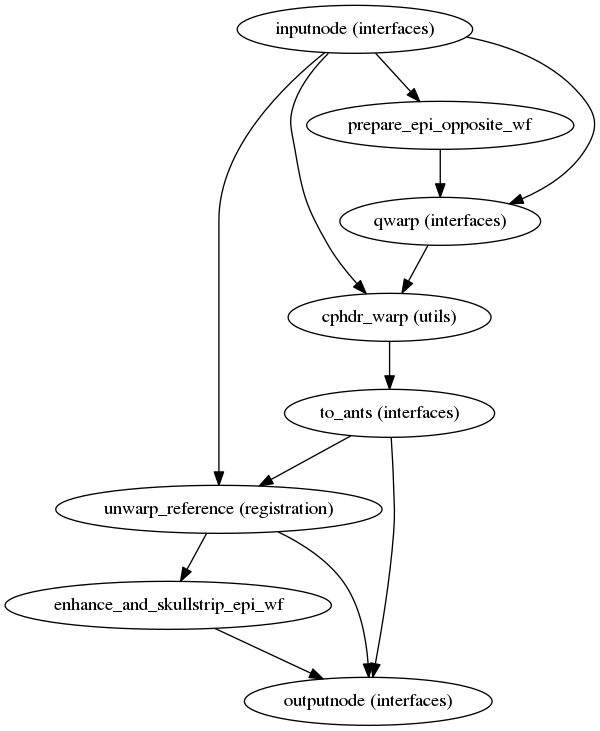
(Source code, png, svg, pdf)
Inputs
- in_reference
- the reference image
- in_reference_brain
- the reference image skullstripped
- in_mask
- a brain mask corresponding to
in_reference - name_source
- not used, kept for signature compatibility with
init_sdc_unwarp_wf
Outputs
- out_reference
- the
in_referenceafter unwarping - out_reference_brain
- the
in_referenceafter unwarping and skullstripping - out_warp
- the corresponding DFM compatible with ANTs
- out_mask
- mask of the unwarped input file
- out_mask_report
- reportlet for the skullstripping
-
fmriprep.workflows.fieldmap.unwarp.init_prepare_epi_wf(ants_nthreads, name='prepare_epi_wf')[source]¶ This workflow takes in a set of EPI files with with the same phase encoding direction and returns a single 3D volume ready to be used in field distortion estimation.
The procedure involves: estimating a robust template using FreeSurfer’s ‘mri_robust_template’, bias field correction using ANTs N4BiasFieldCorrection and AFNI 3dUnifize, skullstripping using FSL BET and AFNI 3dAutomask, and rigid coregistration to the reference using ANTs.
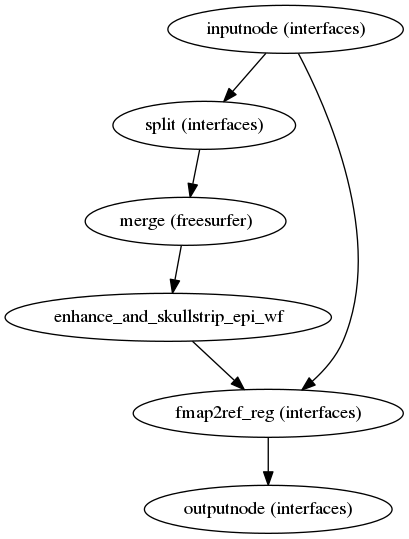
(Source code, png, svg, pdf)
Inputs
- fmaps
- list of 3D or 4D NIfTI images
- ref_brain
- coregistration reference (skullstripped and bias field corrected)
Outputs
- out_file
- single 3D NIfTI file
-
fmriprep.workflows.fieldmap.unwarp.init_sdc_unwarp_wf(reportlets_dir, omp_nthreads, fmap_bspline, fmap_demean, debug, name='sdc_unwarp_wf')[source]¶ This workflow takes in a displacements fieldmap and calculates the corresponding displacements field (in other words, an ANTs-compatible warp file).
It also calculates a new mask for the input dataset that takes into account the distortions. The mask is restricted to the field of view of the fieldmap since outside of it corrections could not be performed.
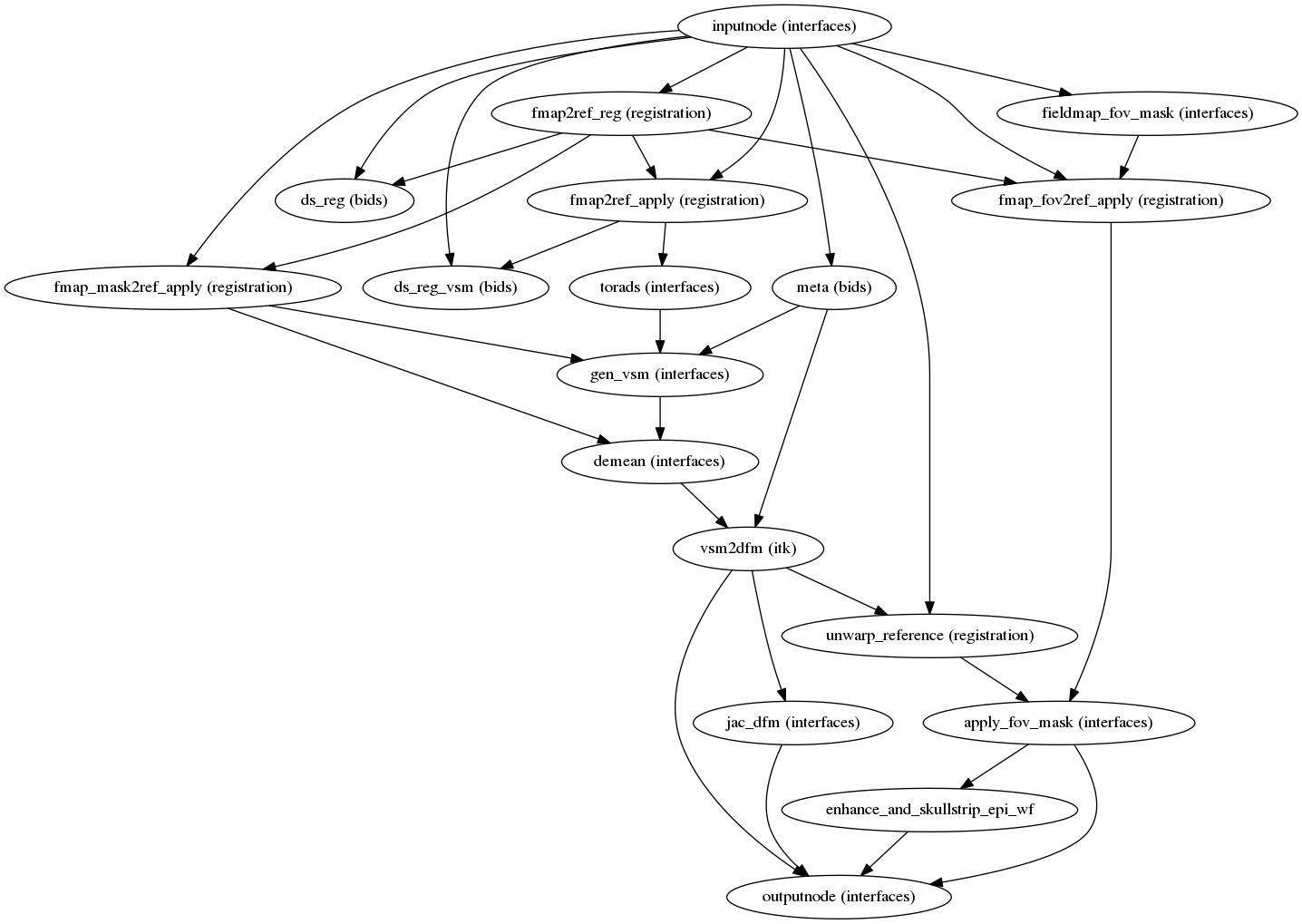
(Source code, png, svg, pdf)
Inputs
- in_reference
- the reference image
- in_mask
- a brain mask corresponding to
in_reference - name_source
- path to the original _bold file being unwarped
- fmap
- the fieldmap in Hz
- fmap_ref
- the reference (anatomical) image corresponding to
fmap - fmap_mask
- a brain mask corresponding to
fmap
Outputs
- out_reference
- the
in_referenceafter unwarping - out_reference_brain
- the
in_referenceafter unwarping and skullstripping - out_warp
- the corresponding DFM compatible with ANTs
- out_jacobian
- the jacobian of the field (for drop-out alleviation)
- out_mask
- mask of the unwarped input file
- out_mask_report
- reportled for the skullstripping